A newly released diagram of all life on Earth, the Tree of Life, contains a whole new branch, full of microbes — which appear to dominate Earth’s biodiversity. How did we miss this?
Since Darwin’s day, scientists have worked to map all of life on a single tree to show how all forms of life on Earth evolved and are related. The DNA sequencing revolution has allowed us to fill in more of the blanks than ever before, greatly accelerating research into biodiversity and Earth’s ecosystems and constantly changing our understanding of life. Last week, our perspective shifted dramatically when researchers unveiled a newly updated Tree of Life showing a whole branch of heretofore-unknown microbes that appear to dominate Earth’s biodiversity.
We asked biodiversity mathematician and TED Fellow Hélène Morlon – who uses computer models and the Tree of Life to better understand the forces that shape evolution – to explain why the new phylogenetic tree is changing the way we view life on Earth.
What is the Tree of Life, and how do you use it in your work?
All known living things are related through common ancestry as diagrammed on a unique tree, which we call the Tree of Life. The leaves of the tree are living organisms, and the branches of the tree represent how these organisms are related to one another. I work with the Tree of Life constantly. It’s the raw material of my research. I use the Tree to understand general principles about the evolution of biodiversity – such as the pace at which species diversify and go extinct – and what factors, such as changes in the environment, affect this pace.
I develop and adjust mathematical models to analyze parts of the Tree of Life, mostly applying this approach to the most familiar parts of it: macroorganisms, like birds and mammals – though I did carry out one study on bacteria, and have another underway in my research group on planktonic diatoms.
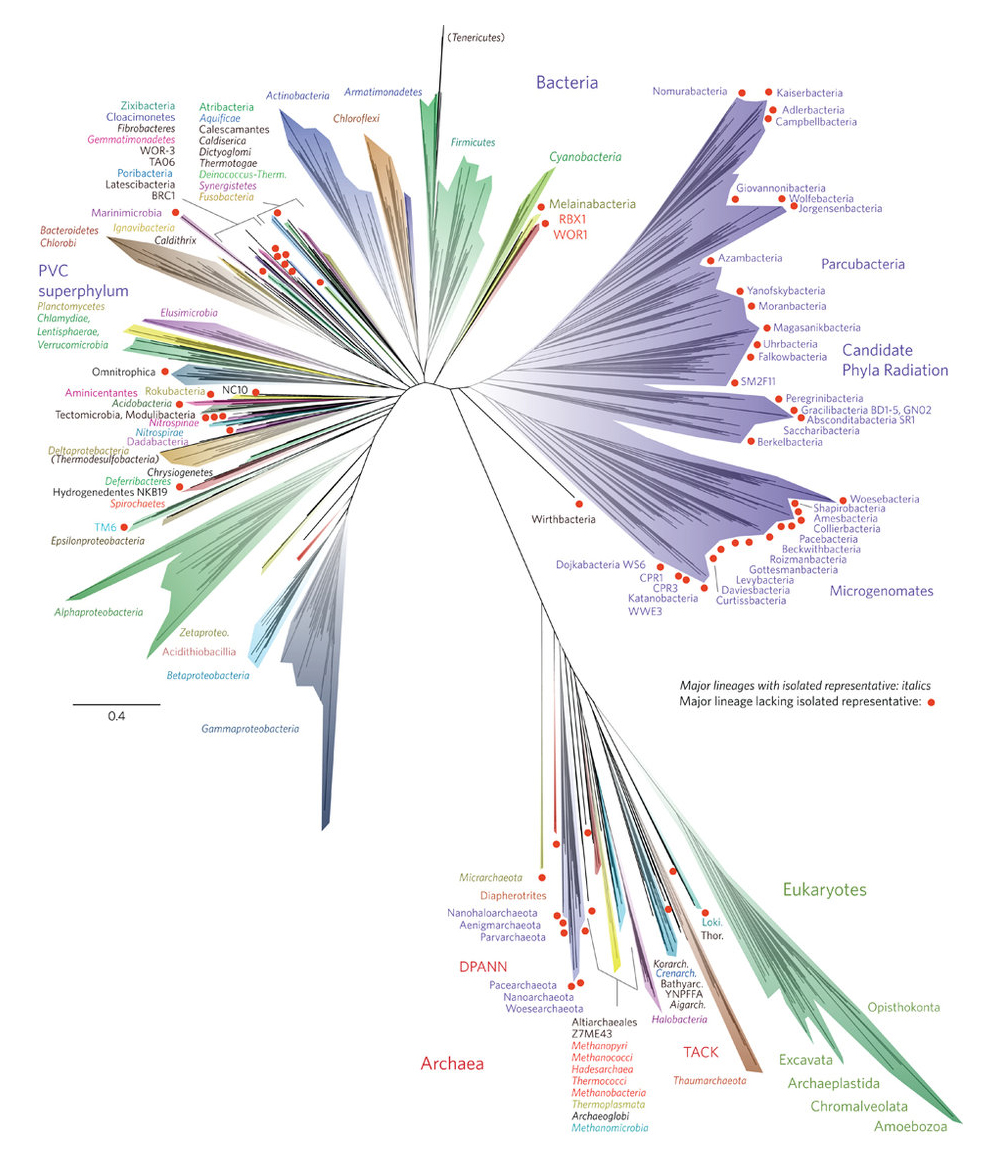
Why does the newly published Tree of Life represent a breakthrough, and what does it tell us?
It’s dramatically reshaping our idea of what the Tree of Life looks like. Since the early 1990s, we’ve thought of the Tree of Life as made up of three major trunks, the three domains of life: Eukarya, animals, plants, fungi and protozoans, the trunk we know the most about; Archaea, comprising single-celled microorganisms; and Bacteria. The newly published Tree more than doubles the size of the Bacteria trunk – and suggests that this domain is subdivided into two. Here’s the mind-blowing part: we’ve been blind to one of these subdomains, which has been coined the “candidate phyla radiation” – yet, it turns out, it contains most of life’s biodiversity. It’s a bit like discovering that we’d missed half of the stars in the Milky Way!
How did scientists not spot this new subdomain before – and how did they manage to discover what was missing?
We didn’t spot it because it doesn’t look like what we thought life should look like. It is a part of life that cannot be cultivated in the lab, and can only be seen via its molecular fingerprint – its DNA. Until now, we thought all bacteria shared a universal fingerprint – the 16S ribosomal RNA gene sequence. In fact, the corresponding gene sequence of bacteria from the “new” bacterial group is quite different, so we often missed it in our traditional surveys.
Instead of relying on 16S, the study’s authors sequenced the whole genetic material from various samples taken from the environment. This metagenomic sequencing technique provides a bunch of tiny gene sequences, and the tricky part is to reassemble them into meaningful genomes. The authors used a series of quite complex bioinformatic algorithms to do this successfully – though we can probably not exclude the possibility that mistakes are being made in this reassembly process.
Why is it significant that this new group of bacteria is so large and contains so much of Earth’s biodiversity?
That this group of bacteria is so dominant means that it’s probably one of the main players in our ecosystems – in terms of nutrient cycling, for example. Yet we know close to nothing about it! It is very distinct from the rest of bacteria: half of the genes are unlike other known genes. We might even find new biological functions that could be useful for designing new drugs or degrading pollutants. As we learn more about this huge group of organisms, we will certainly encounter many new discoveries and surprises.
How much do we not yet know? Is that even a question we can ask?
The fact that we just discovered a megadiverse group that reshapes the overall structure of the Tree of Life demonstrates we know close to nothing about the microbial part of the Tree of Life. This forces us to stay humble about what we do know. This likely won’t be the last time the Tree changes dramatically. We can make useful guesses and estimates, but as this demonstrates, in reality we can have no definite idea about what’s missing. I’m very excited to see progress on the microbial part of the Tree of Life, but there’s still a long way to go. It will not be easy, but to understand the evolutionary history of life on our planet, we need to keep refining the Tree of Life.
The big lesson: we missed an enormous chunk of the Tree of Life because it looked just a bit different from what we assumed life should look like. It’s a beautiful reminder to always question what knowledge we take for granted.